instruction for Res Obj check
Xiangming Cai
2026-01-28
🔍 Introduction
The ResObj Object contains all relavant informations. If you are interested, please read our paper (mentioned on the top) to know the concepts of each matrix shown here. See CreateRes and SlopEstimation for detailed introduction of the structure of ResObj.
💻 Installation
You can install the development version of USERM from GitHub with:
devtools::install_github("xiangmingcai/USERM")Step 1 📈 query avalilable fluorescence list
You can use querySig function to query all available fluorescence.
> head(Sig_info)
id PrimaryName SecondaryName detectors instrument Source Note
1 SCC_Bead_CD4_NFR700 CD4 NFR700 51 Xenith XiangmingCai NA
2 SCC_Bead_CD3_BV510 CD3 BV510 51 Xenith XiangmingCai NA
3 SCC_Bead_CD2_FITC CD2 FITC 51 Xenith XiangmingCai NA
4 SCC_Bead_AF_AF AF AF 51 Xenith XiangmingCai NA
5 SCC_Bead_CD8_BV570 CD8 BV570 51 Xenith XiangmingCai NA
6 SCC_Bead_CD16_BUV805 CD16 BUV805 51 Xenith XiangmingCai NATo check signature of a specific fluroescence, you may
fluor_to_check = Sig_info$id[3]
print(fluor_to_check)
checkSig_linePlot(id = fluor_to_check)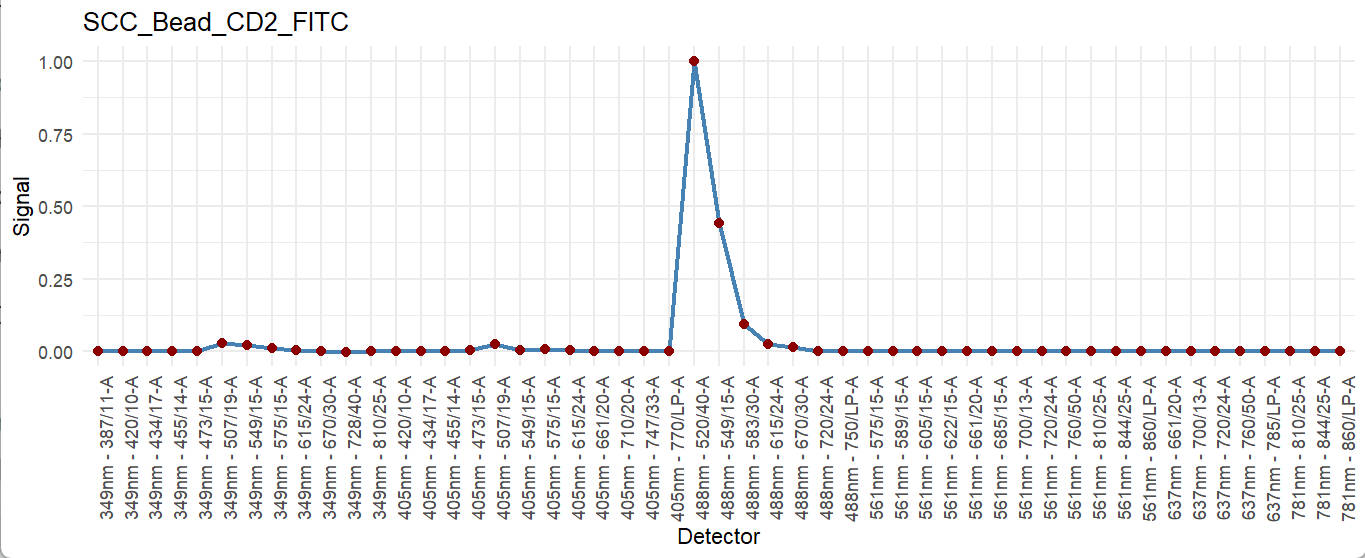
These available fluorescence were preprocessed with parameters extracted for residual model. To access the corresponding residaul model and its parameters, you can use the getRes function.
ResObj = getRes(id = fluor_to_check)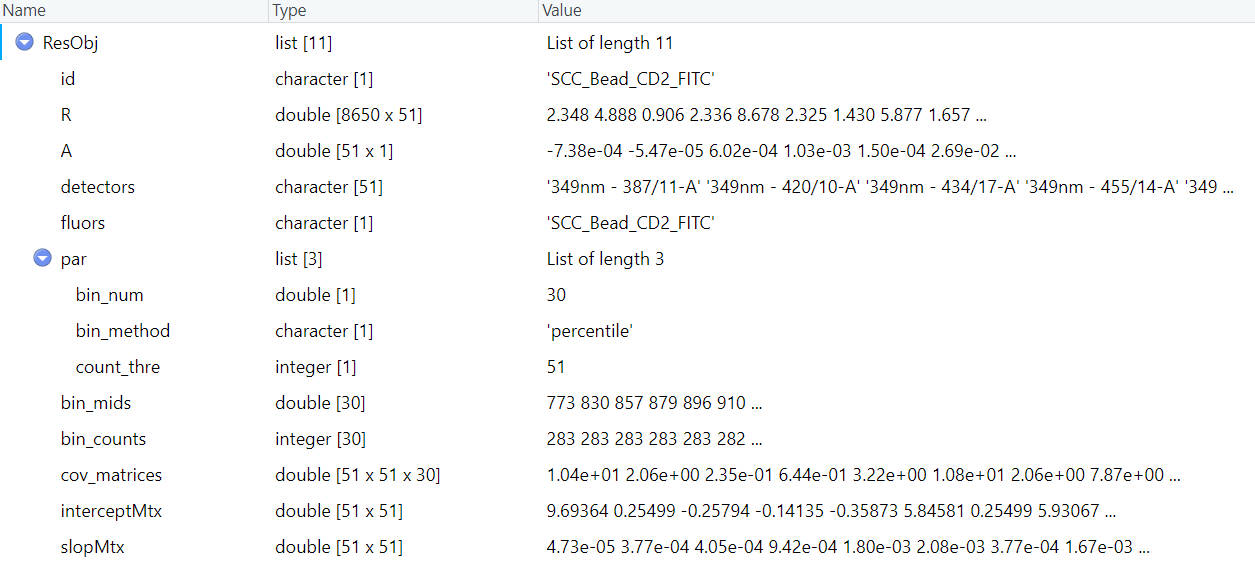
You can easily visaulize all these matrixes, when you are interested in any one of them.
To check the slop matrix:
checkRes_slopMtx(Res = ResObj)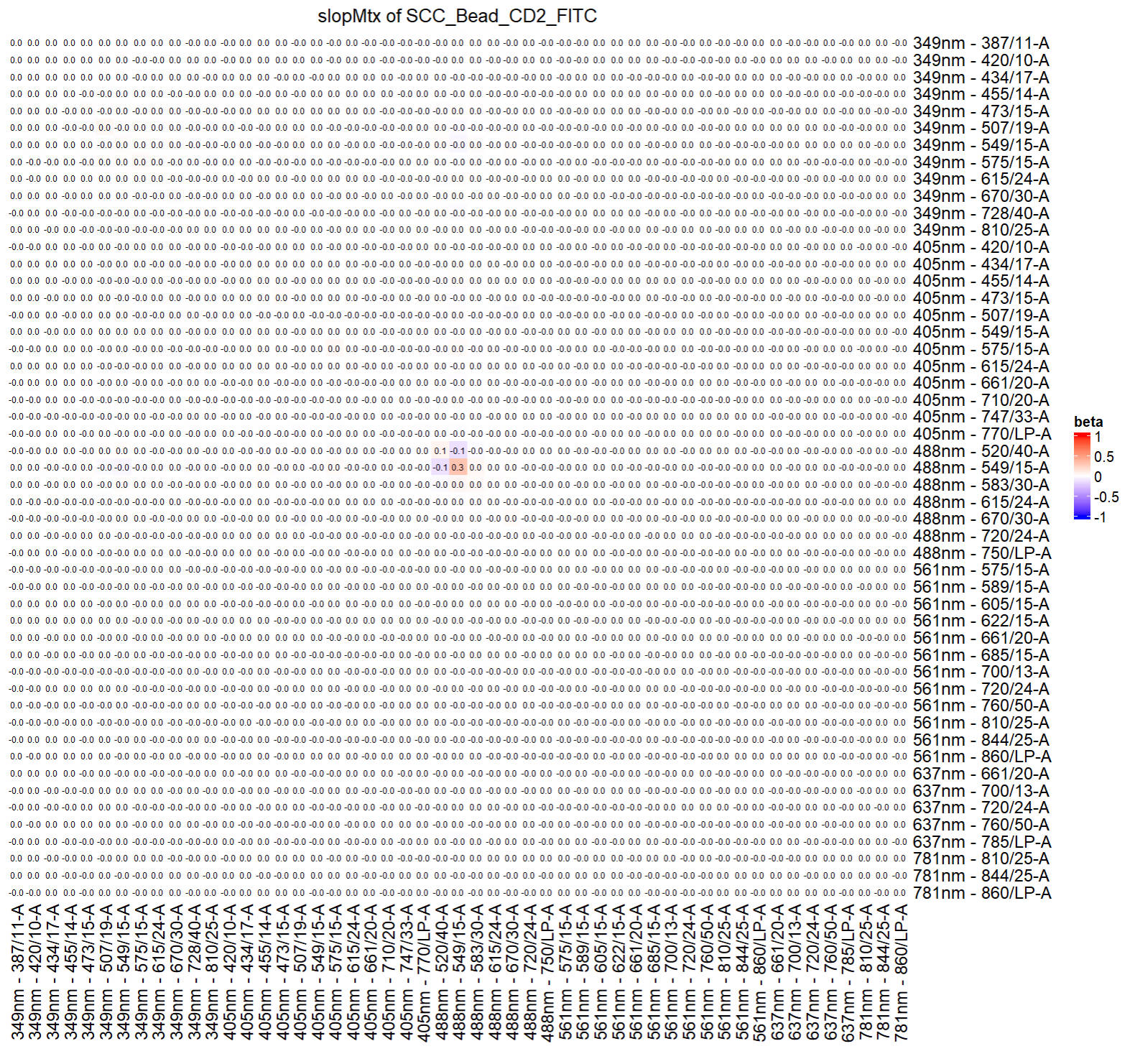
To check the intercept matrix:
checkRes_interceptMtx(Res = ResObj)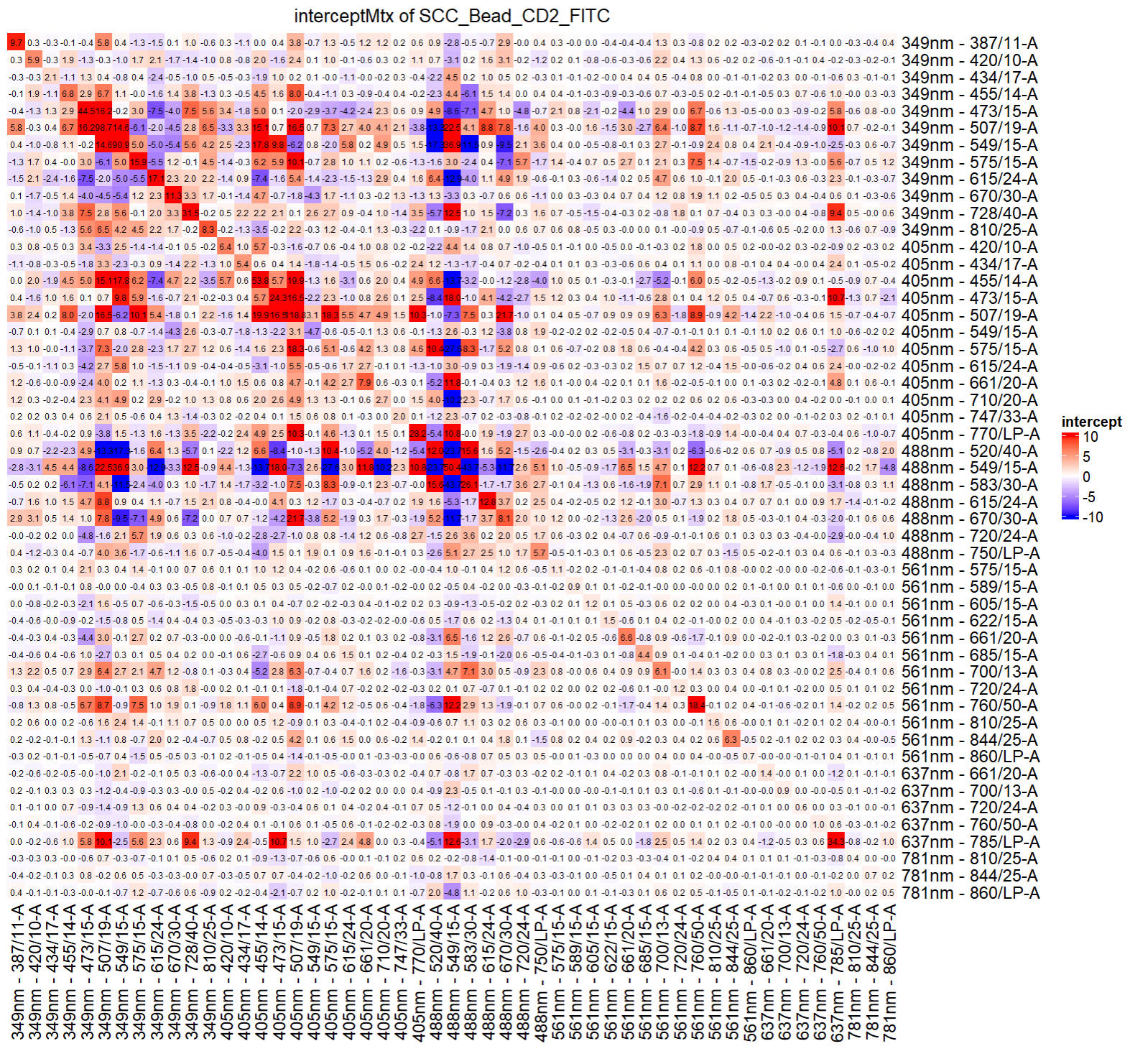
You can check the covariance matrix at a specified fluorescence intensity (bin):
print(ResObj$bin_mids)
checkRes_covMtx(Res = ResObj,bin=3)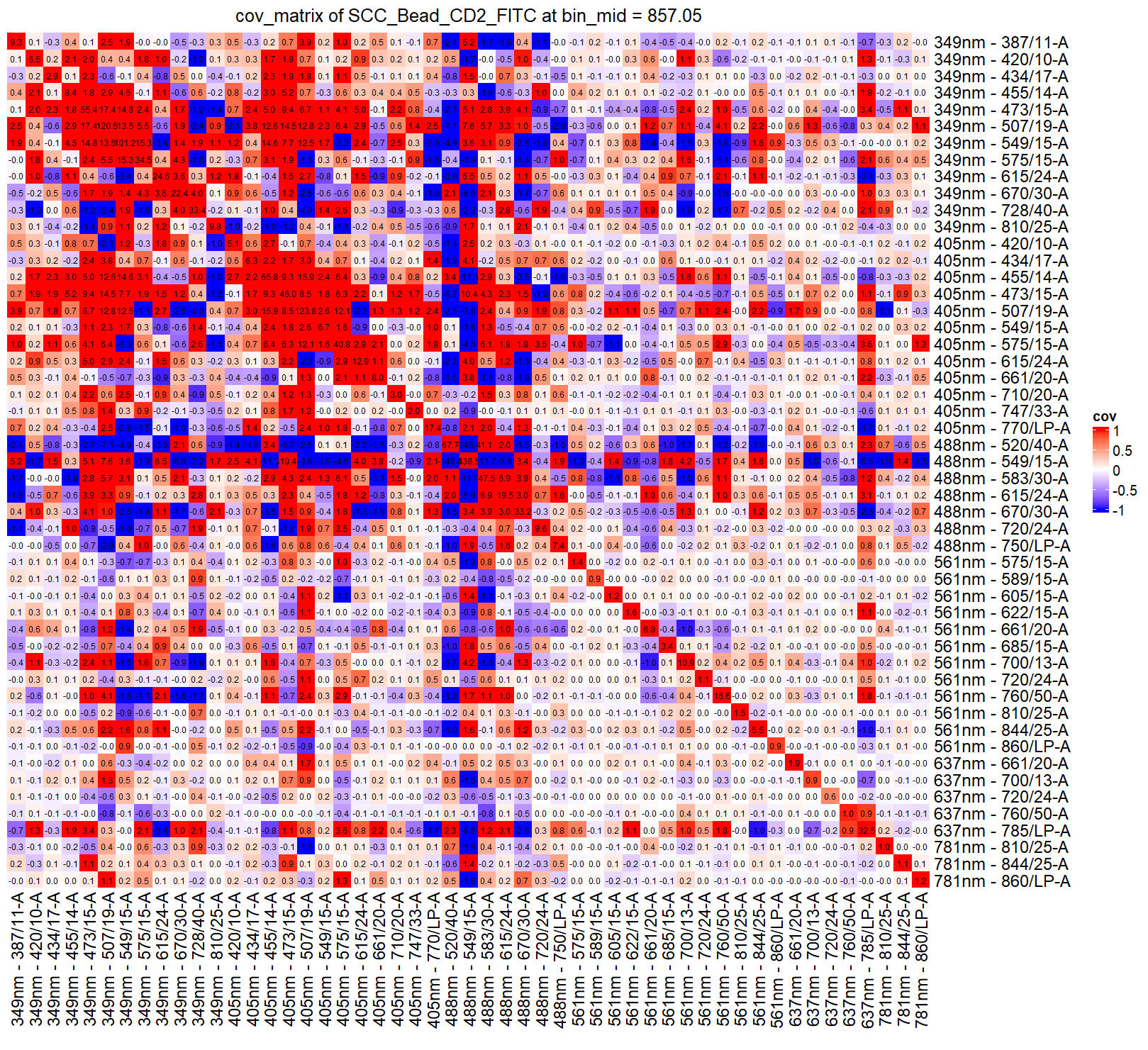
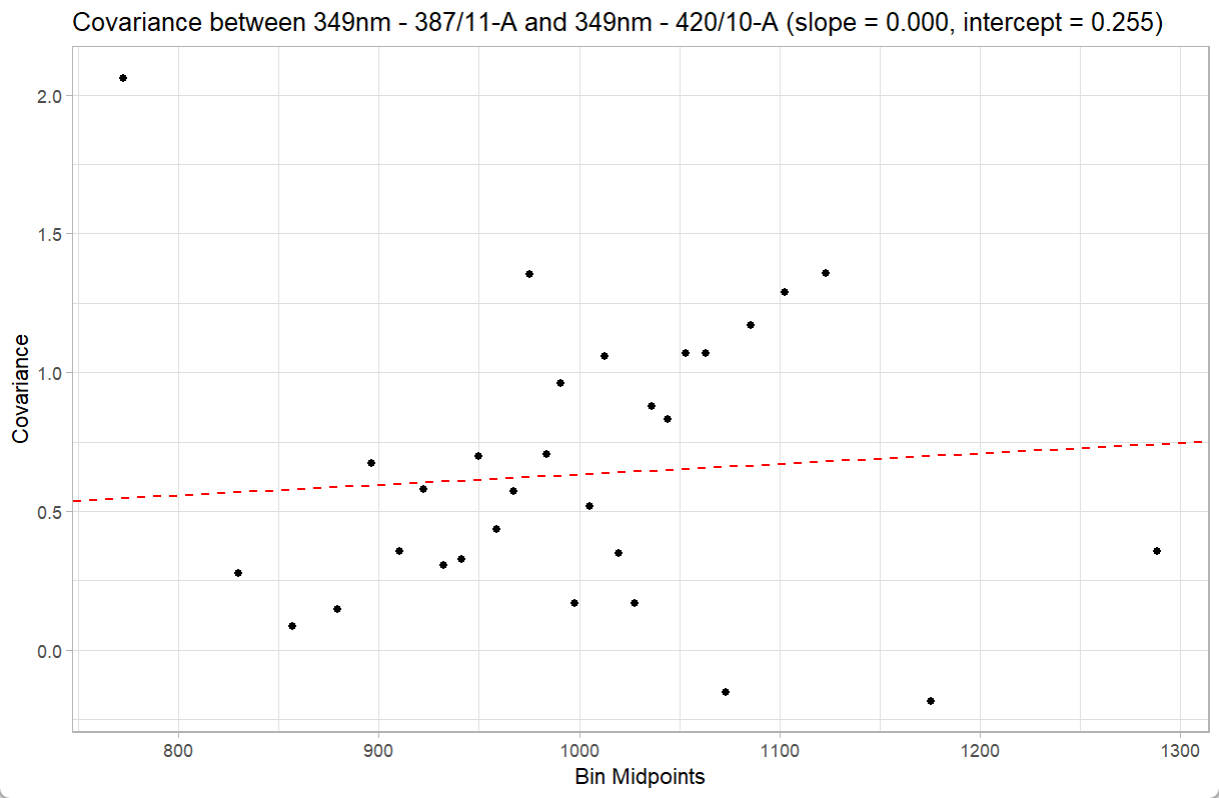
You can even check the linear fitting of covariance versus fluorescence intensity (bin) between two speific detectors:
print(ResObj$detectors)
checkRes_covScatter(Res = ResObj,
detector1 = ResObj$detectors[1],
detector2 = ResObj$detectors[2])